To MiK***: It depends. -> need to order the column according the analyte in your Lab.
Here is the column info I sued @ 2005' Tech.
2: eclipse c18 150 mm * 4.6 mm ,5 um.
1: zorbax-aq 150 mm * 4.6 mm, 3.5 um.
2: zorbax-aq 250 mm * 4.6 mm, 5 um.
4: zorbax-sq 50 mm * 4.6 mm, 5 um.
3: YMC-am-c18 50 mm * 4.6 mm, 5 um.
4: Waters C18 50 mm *2.0mm , 5 um.
LC-MS: Prefer use the zorbax-sq 50 mm *4.6 mm,5 um and Waters C18 50 mm *2.0 mm,5 um, HUGE time saving with Syringe pump.
LC only: eclipse c18 150 mm *4.6 mm and zorbax-sq 50 mm *4.6 mm.
Friday, August 15, 2008
Tuesday, August 12, 2008
DONE! Annual Calibration of Agilent 6890N GC/FID
Update @ Wed.
using the Solution of 0.03% C14, C15 and C16 normal alkanes in hexane.
Then the calibration for Varia** 240 FS Atomic Absorption Spectrometer.
hope get it done NLT this Friday.
Thanks..
using the Solution of 0.03% C14, C15 and C16 normal alkanes in hexane.
Then the calibration for Varia** 240 FS Atomic Absorption Spectrometer.
hope get it done NLT this Friday.
Thanks..
Monday, August 11, 2008
Polydimethylsiloxane (PDMS) Fibers ( non-polar) should be OK for the Flavor Components (2)
To Je***: Based on the searching on my NIST/EPA/NIH 2005 Mass Library, I got some crabs :>)  Peak @ 21.36 min -> 2,6-di-tert-butyl-4-sec-butylphenol
Peak @ 21.36 min -> 2,6-di-tert-butyl-4-sec-butylphenol
 Peak @ 21.36 min -> 2,6-di-tert-butyl-4-sec-butylphenol
Peak @ 21.36 min -> 2,6-di-tert-butyl-4-sec-butylphenolSo please DO NOT trust this Mass Library Searching. will update when I find something new.
Update@Aug-12-2008
Maybe PDMS works. Also it maybe work on the acids, alcohol, aldehydes (ketone). However, that is not a good one. -> “the Supelco 57301 the fiber ( 100 um) PDMS” has better SPE affinity to the high molecular weight non-polar compounds, that maybe the reason you got so many unknown peak ( high molecular compounds).
So if your analyte is low molecular polar acid, alcohol, aldehydes (ketone), recommend the SPME fiber assembly, Carbpwax-Polyethyele (PEG), Supelco#57355-U, it should be good to go for the polar low molecular compounds.
Polydimethylsiloxane (PDMS) Fibers ( non-polar) should be OK for the Flavor Components (1)
To: Je***: Polydimethylsiloxane (PDMS) ( non-polar) should be Good.  What is the thickness of matrix active group (polydimethylsiloxane coating)? using 100μm coating? that 100 μm coating thickness is recommended for low molecular weight or non-polar high molecular weight compounds. Or equivalent with the one in the Your Reference Literature?
What is the thickness of matrix active group (polydimethylsiloxane coating)? using 100μm coating? that 100 μm coating thickness is recommended for low molecular weight or non-polar high molecular weight compounds. Or equivalent with the one in the Your Reference Literature?
If want to obtain a decent result, need to pay attention to the optimization of the SPME method with PDMS: Carry-over effect / Position of the fiber in the GC injector/ Extraction conditions, such as the duration of absorption and desorption, temperature of extraction, and the polarity and structure of the fiber. (Here is the basic SPE info:
http://www.sigmaaldrich.com/Brands/Supelco_Home/Spotlights/SPME_central.html)
If use the 30 μm or a 7 μm PDMS fiber, could extract the non-polar semivolatiles or large molecular weight compounds more effectively.
So my suggestion is: try to do more research on that papers -> find out the right PDMS fiber they used. (100 μm coating or 30 μm or a 7 μm)?
Reference from Varia**: https://www.varianinc.com/media/sci/apps/spme04.pdf
 What is the thickness of matrix active group (polydimethylsiloxane coating)? using 100μm coating? that 100 μm coating thickness is recommended for low molecular weight or non-polar high molecular weight compounds. Or equivalent with the one in the Your Reference Literature?
What is the thickness of matrix active group (polydimethylsiloxane coating)? using 100μm coating? that 100 μm coating thickness is recommended for low molecular weight or non-polar high molecular weight compounds. Or equivalent with the one in the Your Reference Literature?If want to obtain a decent result, need to pay attention to the optimization of the SPME method with PDMS: Carry-over effect / Position of the fiber in the GC injector/ Extraction conditions, such as the duration of absorption and desorption, temperature of extraction, and the polarity and structure of the fiber. (Here is the basic SPE info:
http://www.sigmaaldrich.com/Brands/Supelco_Home/Spotlights/SPME_central.html)
If use the 30 μm or a 7 μm PDMS fiber, could extract the non-polar semivolatiles or large molecular weight compounds more effectively.
So my suggestion is: try to do more research on that papers -> find out the right PDMS fiber they used. (100 μm coating or 30 μm or a 7 μm)?
Reference from Varia**: https://www.varianinc.com/media/sci/apps/spme04.pdf
Saturday, August 9, 2008
How to Calibrate the Uncoded Hollow Cathode Lamps for 240 FS Atomic Absorption Spectrometer
To HJH: Yes, I agreed with your Lab manager -> It is NOT necessary to buy Coded (Certified) Hollow Cathode Lamps for your 220 FS AAS, however, he might neglect to give you some basic explanations.
 For Varia** AAS, I am NOT a Technical Expert, but according my 3-years experiences of Varia*: Coded Hollow Cathode Lamps is only good for the automatic lamp recognition SpectrAA software. So I prefer the Lower priced uncoded Hollow Cathode Lamps.
For Varia** AAS, I am NOT a Technical Expert, but according my 3-years experiences of Varia*: Coded Hollow Cathode Lamps is only good for the automatic lamp recognition SpectrAA software. So I prefer the Lower priced uncoded Hollow Cathode Lamps.
But, always remember that you need to Calibrate the Uncoded Hollow Cathode Lamps before the 1st analysis. 1-3 step is OK.

 For Varia** AAS, I am NOT a Technical Expert, but according my 3-years experiences of Varia*: Coded Hollow Cathode Lamps is only good for the automatic lamp recognition SpectrAA software. So I prefer the Lower priced uncoded Hollow Cathode Lamps.
For Varia** AAS, I am NOT a Technical Expert, but according my 3-years experiences of Varia*: Coded Hollow Cathode Lamps is only good for the automatic lamp recognition SpectrAA software. So I prefer the Lower priced uncoded Hollow Cathode Lamps.But, always remember that you need to Calibrate the Uncoded Hollow Cathode Lamps before the 1st analysis. 1-3 step is OK.
Determination of Ractopamine in Feedstuffs
To Mik**, the reason why failed in the assay of Ractopamine lies on the sample pre-treatment. Here are some information from my note.
(1) Transfer 3.0 g of feedstuff sample into 25 mL solution of 2% Ammonia in MeOH and place onto Vortex 1 min, then sonicate and swirl 5 min. Pipet 10 mL of upper clear solution and transfer to a 1 ml flask, make the dryness @ 50 C by Rotatory Evaporator.
(2) Using 2 mL of MeOH and 2 mL of n-hexane to dissolve the residue, discard the upper solution, and repeat the above procedure twice. Collect the 10 mL of MeOH portion and blow with N2 to dryness @ 45℃. Dissolve the residue again with 5 mL of Ethyl Acetate, add some anhydrous sodium sulfate, and centrifuge the solution@5000rpm for 3 min.
(3) Apply Polystyrene SPE column purify the analyte: (a) Column solvation: apply 5 mL of Ethyl Acetate and 3 mL of Acetonitrile/Ethyl Acetate (1:1). (b) Interference elution: Elute Analyte by using 10 mL of MeOH/Ethyl Acetate (1:1), collect the elute. (c) Concentrate: Under 40 degree C, evaporate on a stream bath to dryness (or with Nitrogen blowing Instrument). Mixing with 0.1 mL BSTFA + 1%TMCS by using Vortex for 1 min, Heat in oven 80 C for 1 hr. Blow with N2 to dryness by Nitrogen blowing Instrument. Bring to the volume with 0.2 mL of toluene for GC-MS analysis.
 GC Column: 5% Phenyl-Methylpolysiloxane 30 m×0.25 mm I.D.×0.25μm.
GC Column: 5% Phenyl-Methylpolysiloxane 30 m×0.25 mm I.D.×0.25μm.
Inlet Temp: 300℃.
Oven: Initial 150℃ (3 min), 10℃/min to 230℃ (10min), 20℃/min to 280℃ (10min).
Carrier Gas: Helium(99.999%)1.0mL/min
Injection: 1.0µL, pulsed splitless, 50 ng on-cloumn per Inj.
EI Temp: 230℃.
MSD Interface:280℃.
EM voltage: 1506V.
Scan Rage: 30~550U.
SIM (m/z): 267、250、502,Quantitative ion 250.
Solvent Delay:5 min. ( I can not forget the accident we made @ Tech Dec-2005. How about you?).
Please let me know if this sample pre-treatment method works.
BTW, How do you valuate the Beijing 2008 Olympic Game Opening ceremony? for me, feel humble in the front of 5000-years China History.
(1) Transfer 3.0 g of feedstuff sample into 25 mL solution of 2% Ammonia in MeOH and place onto Vortex 1 min, then sonicate and swirl 5 min. Pipet 10 mL of upper clear solution and transfer to a 1 ml flask, make the dryness @ 50 C by Rotatory Evaporator.
(2) Using 2 mL of MeOH and 2 mL of n-hexane to dissolve the residue, discard the upper solution, and repeat the above procedure twice. Collect the 10 mL of MeOH portion and blow with N2 to dryness @ 45℃. Dissolve the residue again with 5 mL of Ethyl Acetate, add some anhydrous sodium sulfate, and centrifuge the solution@5000rpm for 3 min.
(3) Apply Polystyrene SPE column purify the analyte: (a) Column solvation: apply 5 mL of Ethyl Acetate and 3 mL of Acetonitrile/Ethyl Acetate (1:1). (b) Interference elution: Elute Analyte by using 10 mL of MeOH/Ethyl Acetate (1:1), collect the elute. (c) Concentrate: Under 40 degree C, evaporate on a stream bath to dryness (or with Nitrogen blowing Instrument). Mixing with 0.1 mL BSTFA + 1%TMCS by using Vortex for 1 min, Heat in oven 80 C for 1 hr. Blow with N2 to dryness by Nitrogen blowing Instrument. Bring to the volume with 0.2 mL of toluene for GC-MS analysis.
 GC Column: 5% Phenyl-Methylpolysiloxane 30 m×0.25 mm I.D.×0.25μm.
GC Column: 5% Phenyl-Methylpolysiloxane 30 m×0.25 mm I.D.×0.25μm.Inlet Temp: 300℃.
Oven: Initial 150℃ (3 min), 10℃/min to 230℃ (10min), 20℃/min to 280℃ (10min).
Carrier Gas: Helium(99.999%)1.0mL/min
Injection: 1.0µL, pulsed splitless, 50 ng on-cloumn per Inj.
EI Temp: 230℃.
MSD Interface:280℃.
EM voltage: 1506V.
Scan Rage: 30~550U.
SIM (m/z): 267、250、502,Quantitative ion 250.
Solvent Delay:5 min. ( I can not forget the accident we made @ Tech Dec-2005. How about you?).
Please let me know if this sample pre-treatment method works.
BTW, How do you valuate the Beijing 2008 Olympic Game Opening ceremony? for me, feel humble in the front of 5000-years China History.
Monday, August 4, 2008
Basic Operation Procedure of Agilent 6890N-5975 MSD (6)
Quantitative Analysis
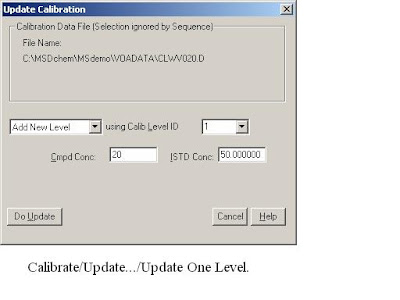
Just like the LC-MS ChemStation Software, also need establish a new calibration table:-> calibration /Setup calibration database. (Here are four Quantitative Methods : AREA%,NORM%, ESTD, ISTD.) The different betw GC and GC-MS is that: Need to Qualtilative before Quantitative Analysis.Load the Method and signals, Select->the Calibrate/Set Up Quantitation
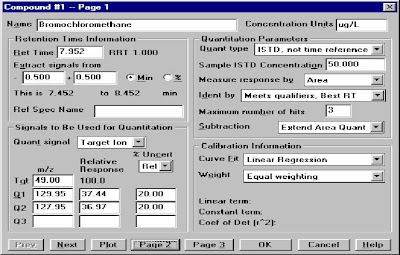
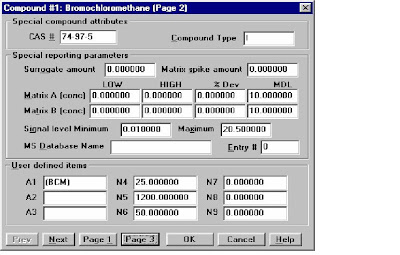
 Quantitation Database Globals:Type the Calibration Title, Default Multiplier, Default Sample Concentration, Reference Window/Non-Reference Window, Reference window for Internal STD,Correlation Window(For Scan: 0.05, For SIM:0.025)
Quantitation Database Globals:Type the Calibration Title, Default Multiplier, Default Sample Concentration, Reference Window/Non-Reference Window, Reference window for Internal STD,Correlation Window(For Scan: 0.05, For SIM:0.025)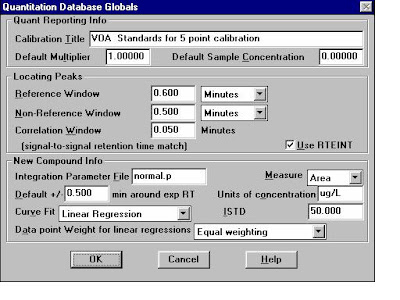
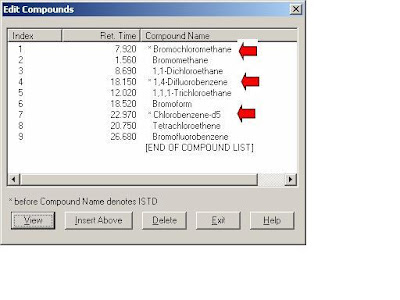
 Double click the Right button, type the chemical name. If it's a internal STD, Select the Internal icon. Move the “+” to select the targeted ions,click the Left/Right button at the same time. DO NOt forget "Save".
Double click the Right button, type the chemical name. If it's a internal STD, Select the Internal icon. Move the “+” to select the targeted ions,click the Left/Right button at the same time. DO NOt forget "Save".Sunday, August 3, 2008
Why the Molecular Weight of the analyte is 335.31?
To Xue*, maybe It's right that your client claimed Molecular Weight is 335.31, NOT 339 that you obtained from NegESI.
C15H17O6N3-Calculated Exact Mass: 12.0000 (15C) + 1.0078 (17H) + 14.0031 (3N) + 15.9949 (6O) =335.30925.
According to the Nitrogen Rule, the chemical structure contains an odd number of nitrogen atoms (3N), indicating an odd nominal mass. So [M+H]+, [M-H]- the molecular ion peak should be an even number. Under negative ESI mode, 334 maybe the [M-H]-. Also under Positive ESI Mode, there are peaks of 336[M+H]+, 358[M+Na]+,371[M+K]+. So the conclusion is 339 could not be [M-H]-.
Also under Positive ESI Mode, there are peaks of 336[M+H]+, 358[M+Na]+,371[M+K]+. So the conclusion is 339 could not be [M-H]-. In practical of NegESI, it is difficult to explain the molecualr ion is the peak of 339: How could it disassociate a Fragments ion of 5 to obtain a Fragments ion of 334. Also, please double check the Mass Spectrum of Positive ESI, there is not Fragments ion of 341. Also confirm that the irrational of molecular ion of 339.
In practical of NegESI, it is difficult to explain the molecualr ion is the peak of 339: How could it disassociate a Fragments ion of 5 to obtain a Fragments ion of 334. Also, please double check the Mass Spectrum of Positive ESI, there is not Fragments ion of 341. Also confirm that the irrational of molecular ion of 339.
As for the fragment peak of 339, that maybe is from the system contamination or the impurity in the analyte. That is the reason why I asked you obtain a positive ESI Mass@ Friday.
C15H17O6N3-Calculated Exact Mass: 12.0000 (15C) + 1.0078 (17H) + 14.0031 (3N) + 15.9949 (6O) =335.30925.
According to the Nitrogen Rule, the chemical structure contains an odd number of nitrogen atoms (3N), indicating an odd nominal mass. So [M+H]+, [M-H]- the molecular ion peak should be an even number. Under negative ESI mode, 334 maybe the [M-H]-.
 Also under Positive ESI Mode, there are peaks of 336[M+H]+, 358[M+Na]+,371[M+K]+. So the conclusion is 339 could not be [M-H]-.
Also under Positive ESI Mode, there are peaks of 336[M+H]+, 358[M+Na]+,371[M+K]+. So the conclusion is 339 could not be [M-H]-. In practical of NegESI, it is difficult to explain the molecualr ion is the peak of 339: How could it disassociate a Fragments ion of 5 to obtain a Fragments ion of 334. Also, please double check the Mass Spectrum of Positive ESI, there is not Fragments ion of 341. Also confirm that the irrational of molecular ion of 339.
In practical of NegESI, it is difficult to explain the molecualr ion is the peak of 339: How could it disassociate a Fragments ion of 5 to obtain a Fragments ion of 334. Also, please double check the Mass Spectrum of Positive ESI, there is not Fragments ion of 341. Also confirm that the irrational of molecular ion of 339.As for the fragment peak of 339, that maybe is from the system contamination or the impurity in the analyte. That is the reason why I asked you obtain a positive ESI Mass@ Friday.
Negative ESI Mode should be OK for Flavonoid
Question from jumn*** Why ESI mode doesn't Work for Flavonoids? According my Tech lab Notebook(#2006-09), my answer to your question is: 100% Negative ESI Mode should be Good for Flavonoids.
According my Tech lab Notebook(#2006-09), my answer to your question is: 100% Negative ESI Mode should be Good for Flavonoids.
 According my Tech lab Notebook(#2006-09), my answer to your question is: 100% Negative ESI Mode should be Good for Flavonoids.
According my Tech lab Notebook(#2006-09), my answer to your question is: 100% Negative ESI Mode should be Good for Flavonoids.Your analyte is belong to Flavonoid family. {Molecular structure of the flavone backbone (2-phenyl-1,4-benzopyrone)}. It contains lot of Phenols, so technically it should be good under Negative ESI mode.
(1) Try to use H2O-MeOH as mobile phase, the Injection Conc. is 1ug/ml.
If above method doesn't work,
(2) Method Development: under Negative ESI mode, direct injection sample through syringe pump. (Use syringe pump, easy tuning & time saving, but need to pay attention on the cleaning of capillary after tuning or could try the six-valve direct injection if feel confident, could obtain a good baseline with min background, time costing:> )
Add some Ammonia to enhance the ion compounds or 5mM of ammonium acetate should be fine. Optimize spray-voltage, voltage plate, capillary voltage/Temp, shealth gas, aux gas, flow rate.
The starting analyte Injection Conc. is from 1 ppm.
If you still have any problem, highly recommend you talk to your BOSS: Double check that is the right "Flavonoid" or the purity of Flavonoid.
Try to use LC to testify your suspect,
Column:2.1 X 50 mm, 5 µm , 100 Å , C18
Temp: 40℃
Mobile: A (H2O) with B (MeOH) 30-70% gradient elute in 5 min.
Flow rate: 1 ml/min.
Wavelength: 340 nm.
if there are multi-peaks in the chromatograms, then maybe explain the problems in NegESI mode. Always talk to your manager first, Don't let him to be the last ONE surprised.
Please update me if have something new, thanks.
Reference Paper: JOURNAL OF MASS SPECTROMETRY, 2004; 39: 312–321: LC-DAD-ESI Tandem MassSpect.
Wednesday, July 30, 2008
Correcting Peak Tailing Problems in Reversed Phase HPLC
Peak tailing in reversed phase HPLC continues to be a common complaint. It is particularly prevalent when separating basic compounds and, therefore, a source of constant problems to those analyzing pharmaceutical compounds by HPLC.
(1) Cause: Sample solvent stronger than the mobile phase.
(1) Cure: Dissolve sample in mobile phase or at least reduce the strength of the sample solvent as much as possible.
(2) Cause: Peak Tailing Caused by Sample Mass Overload.
(2) Cure: Reduce the amount (mass) of sample injected, choose injection size for different column configurations.
(3) Cause: Peak Tailing Caused by Stationary Phase Silanol Interactions with Amines.
(3) Cure: Reduce mobile phase pH to < 3.0.Increase mobile phase ionic strength. 25mM to 50mM recommended.Add a competing amine to the mobile phase.10 mM TEA is usually sufficient.Select a stationary phase with a lower silanol activity.
(4) Cause: Peak Tailing Caused by Adsorption of Acidic Compounds onto Silica.
(4) Cure: To correct peak tailing in these cases increase the salt concentration of the mobile phase to suppress secondary interactions, reduce the mobile phase pH to protonate silanols and solutes and, if necessary, add a competing acid to the mobile phase.
(5) Cause: Peak Tailing Caused by a Void in the Column's Packing Bed.
(5) Cure: The best cure is to replace it. This saves time, money, and frustration.or call me if DON'T have the Lexus.
The Final Cure to # 3: Add a competing amine to the mobile phase. Triethylamine (TEA) is commonly added to mobile phases for this purpose. TEA interacts strongly with silanols and inhibits them from interacting with amines in the sample. About 10mM TEA is sufficient for most applications.
Ace C18 <>
Thanks for MAC-MOD Ana Inc. Tech Report # 07031TR
Subscribe to:
Comments (Atom)